The SARS-CoV-2 pandemic has permeated almost every sphere of human endeavour for the last 18 months and academic research is no different. While we lack the ability to see into the future, mathematical modelling studies have been used throughout the pandemic to: make predictions about the future course of the pandemic, explain SARS-CoV-2 dynamics, evaluate different scenarios or strategies, and to provide an evidence base for trials. Results from these modelling studies have been used to inform government policy and interventions around the world. Naturally, with something that has such a significant effect on people, there has been much discussion, debate and even misinformation about these modelling studies.
We have been working with Martyn Fyles, a Mathematics PhD student at the University of Manchester (funded by the Alan Turing Institute), and Liz Fearon, an associate professor in Epidemiology from the London School of Hygiene and Tropical Medicine. Martyn and Liz developed a simulation model of contact tracing, which can be used to predict and explain the effectiveness of different strategies in reducing the spread of SARS-CoV-2.The idea of modelling studies like this one is to combine existing scientific knowledge and data to provide a description of a system. This can then be used to consider different outcomes when changes are made to the system. This allows researchers to see the effect of various policy interventions. All models must select enough features of the real world to give it useful predictive properties, without being so complex that the model becomes intractable.
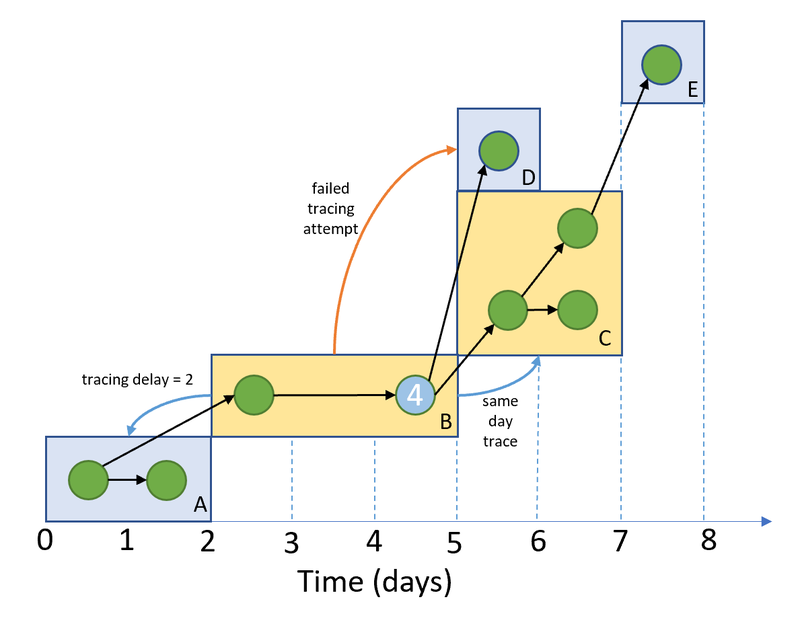
By the beginning of 2021, Martyn and Liz had developed an effective model and were submitting reports to the Scientific Pandemic Influenza Group on Modelling (SPI-M), and SAGE, UK government committees dealing with the pandemic. While the model was working well, it had been developed rapidly and was starting to become difficult to maintain. This is a common stage for many code bases used in research where the code is correct and functional, but becomes more complex and more difficult to maintain and add new features to.
The process of changing code to improve its structure is called refactoring. The idea of refactoring is that the results given by the code should not change, but the structure of the code should change to make it easier to understand, more flexible, and easier to maintain and extend in the future. There was significant controversy in 2020 surrounding the code used by Professor Neil Ferguson in an influential COVID-19 study. The study was based on code which had been in development for over 10 years without significant refactoring, which meant the results were hard to reproduce.
Martyn and Liz’s code certainly hadn’t reached this extreme, but needed some attention. Over the next few months we worked on refactoring the code to make the code more extensible and to make results easier to reproduce. The refactored codebase is now available on GitHub (pre-release version).
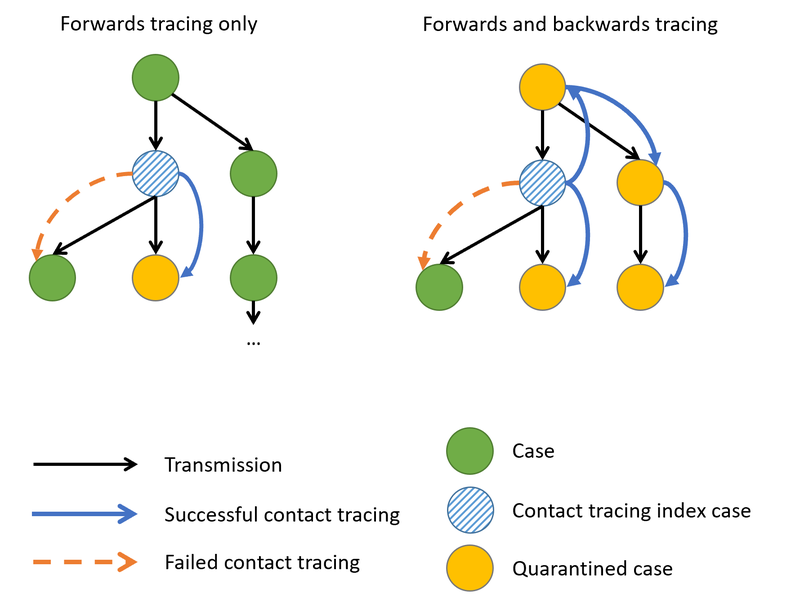
Martyn said “Ann and Peter have done an excellent job refactoring our code base, and I have learnt a lot from working with them. The code is now significantly more modular and extensible, which will allow us to respond to modelling queries with more ease, and without adding maintenance overhead. The refactored code will soon be released with an open source license, allowing it to be used and extended around the world. It is important that we maintain and preserve models, so that they can easily be adapted for future epidemics, and Ann and Peter’s work has made this possible.”
If your research code is getting hard to maintain or you are finding it harder to extend or add new features, perhaps you could benefit from a consultation with the Research IT group. You can ask for a consultation on our Contact page.
This work was supported by UKRI grant MR/V028618/1.